Community powered conda packaging channel conda-forge
2016 DMAC
The problem
The Python scientific community always wanted a package manager that is cross platform, does not require sudo, and lets Python be awesome ;-)
The conda package manager solved that problem, but created a new ones...
Why not wheels?
More about the problem
- How to get software that is not in the
defaultchannel? - How to build the binaries to ensure they are compatible with other systems, and the default channel?
- Which community channels are OK to use? (
anaconda search -t conda gdalreturns 48 options!)
Many choices is not always good!
The default channel does not keep up with the pace of the scientific community and many users/communities channels were created to fill in that gap.
Many choices is not always good!
That led to
- duplication of effort
- recipe fragmentation
- unstable environments when mixing and matching packages from different channels.
The solution
The conda-forge organization was created to be transparent, open, and community led. The goal is to provide a stable source for packages while reducing the effort duplication and recipe fragmentation.
While developed to meet the unique needs of scientific software developers, it is a system that brings features and utilities for the broader developer community.
Some background
- The scientific community adopted conda
- Many channels popped up:
ioos,scitools,bioconda,astropy, etc. - Tons of duplicated effort, fork confusion, bit rot...
See http://wesmckinney.com/blog/conda-forge-centos-moment for more on conda-forge
(Yeah... Wes was quicker than us to write about it!)
But we finally got our post out in time for DMAC!
https://www.continuum.io/blog/developer-blog/community-conda-forge
What do we mean by community?
- The channel is community-governed: open by-weekly meetings, and transparent discussions on GitHub (still working on a CoD)
- Open meeting minutes: https://conda-forge.hackpad.com/
- All the tools used to built the binaries are published and available.
There is a vetting process for submitting, verifying, and storing signed releases.
The conda-forge channel
70102104 contributors432579583 packages (IOOS: 224, SciTools: 27)- Available platforms: Linux-64, Windows-32/64, and OS X
- Use with
conda install -c conda-forge gdalconda config --add channels conda-forgeDistributed vs the centralized model
- Individual repositories (feedstocks)
- Teams of maintainers for each feedstock
- Python scripts as Heroku webservices to tie everything together
Builds
- The recipes are hosted on its own feedstock GitHub
- AppVeyor ➛ Windows
- Travis-CI ➛ OS X
- CircleCI+Docker ➛ Linux (CentOS 5)
Automation
- The point of entry is staged-recipes
- The tooling lives in conda-smithy
- Once the PR is accepted a GitHub team is created based on the maintainers list
- The maintainers have commit rights only to their own recipes
Linter

conda smithy recipe-lintUpdating a feedstock
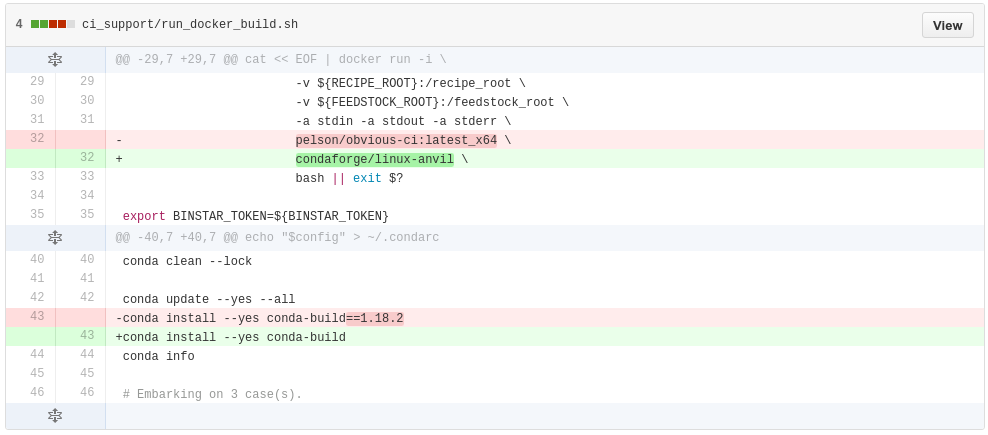
conda smithy rerenderIssues
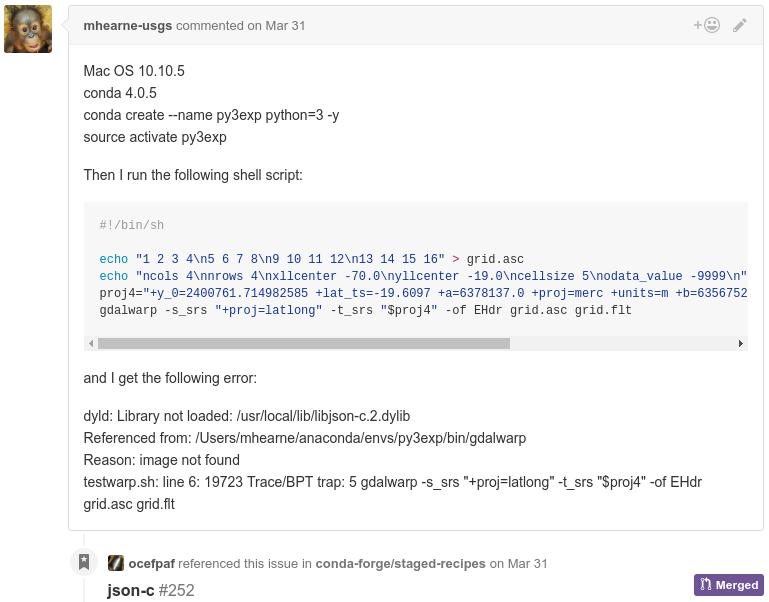
How to contribute?
- Report problems
- Request packages or new releases
- Send PRs adding/fixing packages
Submitting a new recipe
- Fork https://github.com/conda-forge/staged-recipes
- Submit the PR
- The recipe will be built and, once merged, it will trigger the creation of the feedstock repository
- In the feedstock repo, the recipe will be re-built and uploaded to the conda-forge channel on anaconda.org
Updating a recipe
The changes
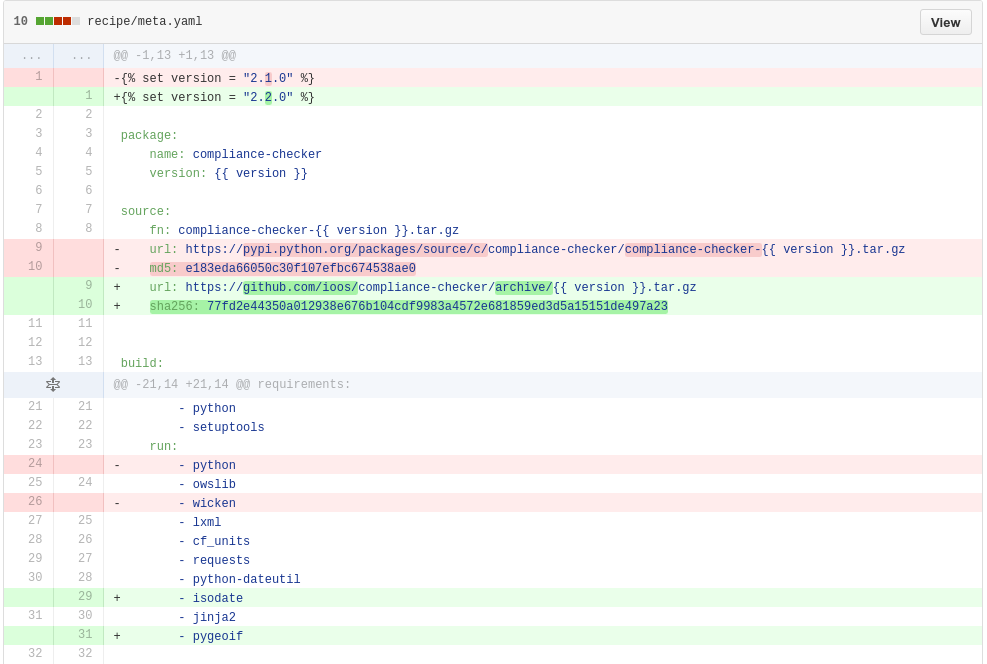
Provenance
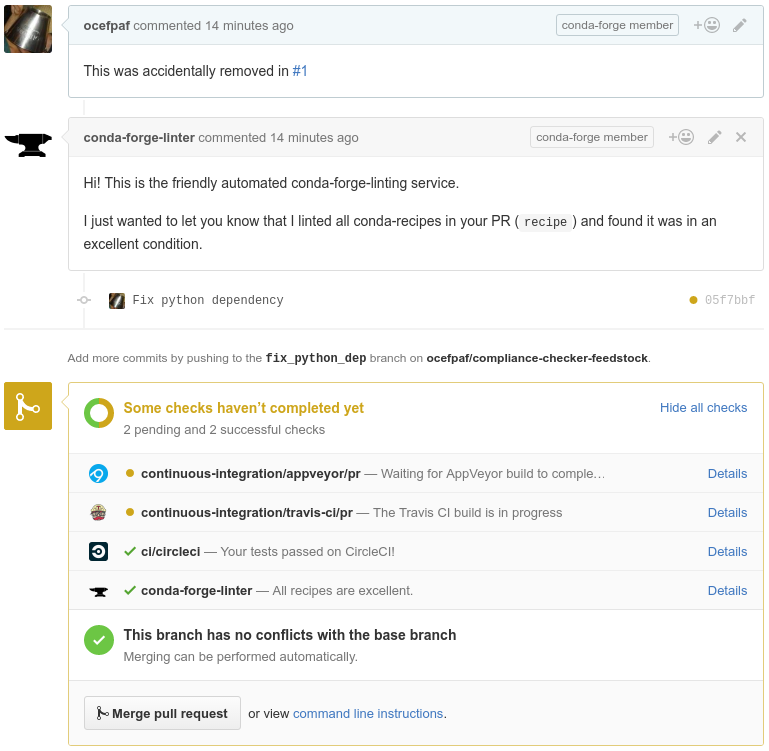
conda-forge outside IOOS